|
max ()) # plot class samples for idx, cl in enumerate ( np. contourf ( xx1, xx2, Z, alpha = 0.4, cmap = cmap ) plt. arange ( x2_min, x2_max, resolution )) Z = classifier. arange ( x1_min, x1_max, resolution ), np. predict ( Z_pca_test ) print ( 'Accuracy (tree_pca): %.2f ' % accuracy_score ( y_pca_test, y_pca_pred ))ĭef plot_decision_regions ( X, y, classifier, test_idx = None, resolution = 0.02 ): # setup marker generator and color map markers = ( 's', 'x', 'o', '^', 'v' ) colors = ( 'red', 'blue', 'lightgreen', 'gray', 'cyan' ) cmap = ListedColormap ( colors ) # plot the decision surface x1_min, x1_max = X. fit ( Z_pca_train, y_pca_train ) y_pca_pred = tree_pca. load ( './output/Z_pca.npy' ) # random_state should be the same as that used to split the Z_forest Z_pca_train, Z_pca_test, y_pca_train, y_pca_test = train_test_split ( Z_pca, y, test_size = 0.3, random_state = 0 ) tree_pca = DecisionTreeClassifier ( criterion = 'entropy', max_depth = 3, random_state = 0 ) tree_pca. predict ( Z_forest_test ) print ( 'Accuracy (tree_forest): %.2f ' % accuracy_score ( y_forest_test, y_forest_pred )) # train a decision tree based on Z_pca # load Z_pca that we have created in our last lab Z_pca = np. 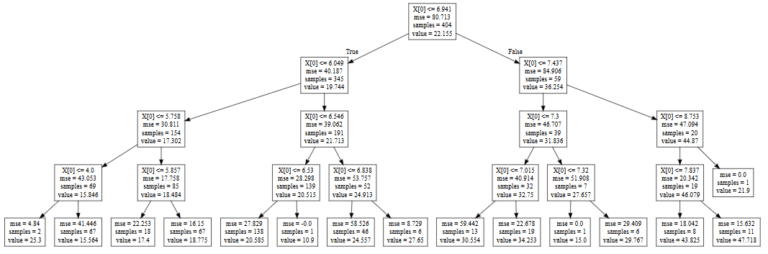
fit ( Z_forest_train, y_forest_train ) y_forest_pred = tree_forest. # train a decision tree based on Z_forest Z_forest_train, Z_forest_test, y_forest_train, y_forest_test = train_test_split ( Z_forest, y, test_size = 0.3, random_state = 0 ) tree_forest = DecisionTreeClassifier ( criterion = 'entropy', max_depth = 3, random_state = 0 ) tree_forest. Another advantage is that the computational cost can be distributed to multiple cores / machines since each tree can grow independently. Generally, the larger the number of trees, the better the performance of the random forest classifier at the expense of an increased computational cost. The only parameter that we need to care about in practice is the number of trees $T$ at step 3. Therefore, we typically don't need to prune the trees in a random forest. Aggregate the predictions made by different trees via the majority vote.Īlthough random forests don't offer the same level of interpretability as decision trees, a big advantage of random forests is that we don't have to worry so much about the depth of trees since the majority vote can "absorb" the noise from individual trees.Repeat the steps 1 to 2 $T$ times to get $T$ trees.Split the node by finding the best cut among the selected features that maximizes the information gain.Randomly select $K$ features without replacement.Grow a decision tree from the bootstrap samples.Randomly draw $M$ bootstrap samples from the training set with replacement. 
The random forest algorithm can be summarized in four simple steps: The idea behind ensemble learning is to combine weak learners to build a more robust model, a strong learner, that has a better generalization performance. Intuitively, a random forest can be considered as an ensemble of decision trees. Random forests have gained huge popularity in applications of machine learning during the last decade due to their good classification performance, scalability, and ease of use. 
Visualize $f^∗$ so we can interpret the meaning of rules.Randomly split the Wine dataset into the training dataset $\mathbb)$$ using the testing dataset.Followings are the steps we are going to perform: Now, consider a classification task defined over the Wine dataset: to predict the type (class label) of a wine (data point) based on its 13 constituents (attributes / variables / features).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
